Overview
The ![]() Find SNVs function can be use to find all single nucleotide variants (SNV) between Samples or between Samples and the target reference sequence(s) of the Task Template(s). The function is available in the comparison table, and extracts the SNVs from the allele sequences that are corresponding to the distance columns (those with green colored headings) of the comparison table.
Find SNVs function can be use to find all single nucleotide variants (SNV) between Samples or between Samples and the target reference sequence(s) of the Task Template(s). The function is available in the comparison table, and extracts the SNVs from the allele sequences that are corresponding to the distance columns (those with green colored headings) of the comparison table.
Starting SNVs Search
When the Find SNVs function is invoked, a handling of missing values dialog window is shown if the table contains any distance columns with missing data. By default all Samples with more than 10% missing targets will be removed from the table. Alternatively or in addition all columns with any missing data can be removed. If no missing data exists, this dialog is skipped.
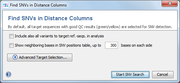
Next another Find SNVs in Distance Columns dialog window opens, that allows to specific the following options before the SNV search is conducted:
- Include also all variants to target ref.-seqs. in analysis
- If this option is enabled, all Sample target sequences are also compared to the ref.-seq. of the targets that are defined in the Task Template(s). Thus all differences to the target ref.-seqs. are also returned as SNV positions, even if all selected Samples exhibit an identical nucleotide at these positions. If the option is not selected, only the positions of SNVs between the selected Samples are returned.
- Show neighboring bases in SNV positions table
- If this option is enabled, the resulting table contains additional columns with neighboring bases next to the SNV position. The export of neighboring bases can be used to design primers (e.g., to confirm by Sanger sequencing the SNVs). A column with neighboring bases contains up to a specified number of bases (default 300) from each side of the SNV position. The SNV position itself is indicated with a lowercase character. This column is followed by a second column with the number of bases that were exported (default 601 or lower, if the SNV position lies near the start or end of a target), and a third column with the SNV position. There are three different types of neighboring bases: exported from the first Sample that contains this SNV, the ref.-seq. of the target, and the seed genome (if applicable also with intergenic region bases) of the Task Template. If the target or the SNV is not found in the first Sample, additional columns from Samples that contain the SNV are inserted next to the first Sample columns.
- Retrieve coverage for SNV positions from stored reads
- If this option is enabled, the resulting table contains additional columns with the coverage at SNV positions for samples where the aligned reads were stored.
- Advanced Target Selection
- This button can be used to select specific targets only. By default, all target sequences with good QC results (green/yellow) are selected for SNV detection.
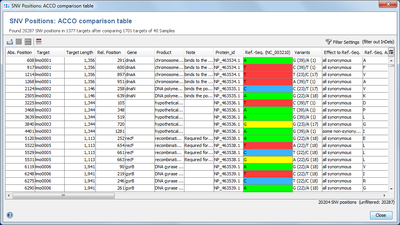
Result of SNV Search
The result of the SNV search is shown in a table, where each row represents a found SNV position. The columns Variants summarizes the bases and aminoacids at this position and the frequencies in the Samples.
The following functions are available via the toolbar:
- Export Table: Export the table to an Excel or CSV file.
- Multiple Alignment: Once a row is selected this command creates a multiple alignment for the target of the selected SNV position for some or all Samples. It is not recommended to do a multiple alignment of more than 100 Samples due to time and especially memory constrains. Therefore, a Sample selection dialog is shown first that allows also to remove Samples from multiple alignment analysis.
- Show/Hide all Sample SNV Columns: By default only the summary columns for the variant bases is shown. Use this function to show all Sample columns.
- Open in Comparison Table: The function can be used to create a Sample-centric comparison table that allows to calculate and visualize distances between Samples based on the nucleotides of the found SNV positions.
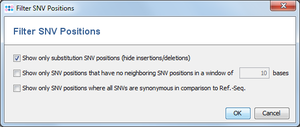
- Filter Settings: The Filter SNV Positions dialog can be used to show only a subset of the SNV positions in the table. Up to three different filter options can be specified:
- Show only substitution SNV positions (hide insertions/deletions)
- This hides all SNV positions that have any insertions or deletions. This option is by default turned on and only shown if there are at all any insertions or deletions present.
- Show only SNV positions that have no neighboring SNV positions in a window of # bases
- This filter searches for neighboring SNV positions within the specified window of bases. Only if two neighboring SNVs are found in at at least one Sample these positions are hidden for all Samples.
- Show only SNV positions where all SNVs are synonymous in comparison to Ref.-Seq.
- This hides all SNV positions that have an non-synonymous change in the amino acid at the SNV position (compared to Ref.-Seq. column). The synonym/non-synonym effect is also shown in the column Effect to Ref.-Seq.
- Show only SNV positions that differ in specific samples
- This filter allows to select two or more samples, and hides all SNV positions that equal in the selected samples.
The table can be resorted by clicking on the column header if it contains less than 5000 SNV rows.