Contents
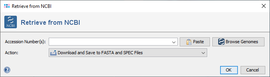
Download by Accession Number(s)
This window allows to download sequences from NCBI GenBank. The button ![]() Paste can be used to get accession numbers from clipboard or from a text file.
Paste can be used to get accession numbers from clipboard or from a text file.
The button ![]() Browse Genomes opens the NCBI GenBank Bacteria Genoms Browser.
Browse Genomes opens the NCBI GenBank Bacteria Genoms Browser.
Download by Browsing
The Ridom Typer NCBI Browser allows to browse two sources:
- NCBI Datasets
- NCBI Genome Reports (legacy)
Note that NCBI Genome Report will return genomes accession numbers (e.g., NC_002695) while a NCBI Datasets search will return assembly accession numbers (e.g., GCF_000008865). A genome accession is an accession number assigned to a single sequence in GenBank while an assembly accession identifies the entire genome assembly, the complete set of sequences that together represent the genome.
![]() Important: As the NCBI prokaryotic genome report is not updated any more, NCBI Genome Report contains only genomes that were released before October 2024. It is therefore recommended to use the NCBI Datasets and the assembly accession numbers.
Important: As the NCBI prokaryotic genome report is not updated any more, NCBI Genome Report contains only genomes that were released before October 2024. It is therefore recommended to use the NCBI Datasets and the assembly accession numbers.
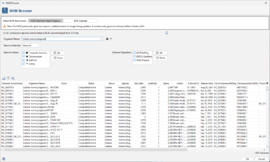
Browse Datasets
The Dataset Browser allows to search within the NCBI Datasets. A web service is used to query the NCBI database. Enter a species name or NCBI taxon ID into the Taxon field and click the Search button. The first 1,000 results returned from NCBI will be displayed. Select rows in the table and click OK to use the accession numbers for an NCBI genome download.
The Search text field, the assembly level selection and the date fields can be used to limit the search results. Use the Reference only checkbox to limit the results to reference genomes. If such limiter(s) are used the whole NCBI Datasets are searched and not just the shown 1,000 results.
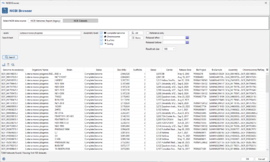
Browse NCBI Genome Report (legacy)
This tab shows a table of the bacterial genomes that are available in the NCBI prokaryotic genome report (up to October 2024). Right-click in the table to open related pages in your Web Browser. Select rows in the table and click OK to use the accession numbers for an NCBI download. The table can be filtered using the fields above:
The field Organism Name can be used filter the complete list of genomes for just one organism. Press cursor key right or the button '![]() ' to auto-complete the typed text and show a list of possible organisms, sorted by the frequency of entries per organism.
' to auto-complete the typed text and show a list of possible organisms, sorted by the frequency of entries per organism.
The field Genome Data Set define which data sequence(s) should be downloaded:
- Genome
- Chromosome
- Plasmids
The field Genome State can be used filter by the state that of the genome data:
- Complete Genome
- Chromosome
- Scaffold
- Contig
The field Genome Repository can be used to select only specific sources for the genomes:
- NCBI RefSeq (NCBI Reference Sequence Database)
- INSDC/GenBank (International Nucleotide Sequence Database Collaboration: Made up of NCBI GenBank, European Nucleotide Archive, and the DNA Data Bank of Japan)
- WGS Projects (Whole genome shotgun sequencing projects at NCBI)
FOR RESEARCH USE ONLY. NOT FOR USE IN CLINICAL DIAGNOSTIC PROCEDURES.