The Picard (citation) library is used in Ridom Typer to process SAM/BAM files. It also provides functions to calculate metrics for validating library construction including the insert size distribution.
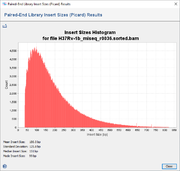
The menu function Tools | Genome Utilities | llumina Insert Sizes (Picard) can be used to analyze the distribution of insert sizes of aligned reads. When invoked the SAM/BAM file that should be analyzed must be selected. After the file reading has finished, a windows is opened showing a histogram for the distribution of insert sizes for all reads. Below the histogram the statistical values for mean, standard deviation, median, and mode are shown.
![]() Hint: This function expects a SAM/BAM file with aligned reads. To analyze the insert size of unaligned reads (i.e, FASTQ files), they must first be mapped to a reference.
Hint: This function expects a SAM/BAM file with aligned reads. To analyze the insert size of unaligned reads (i.e, FASTQ files), they must first be mapped to a reference.
FOR RESEARCH USE ONLY. NOT FOR USE IN CLINICAL DIAGNOSTIC PROCEDURES.