Contents
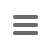
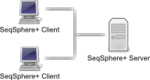
Client/server architecture
The client/server architecture allows distributed workgroups to cooperate via intra- or internet. Multiple users can access the data from various computers. Access rights and privileges can be configured to restrict access for specific users or groups.
The SeqSphere+ software consists of a client application and a server application:
- The SeqSphere+ Server has an integrated SQL database and can be connected by SeqSphere+ clients via LAN or internet.
- The SeqSphere+ Client is a desktop application that must be installed on your local computer.
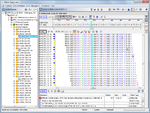
DNA re-sequencing editor
WGS data: Assemble data using the integrated de-novo Assembler Velvet or Spades, or the BWA reference mapper. Edit and analyze assembled DNA sequences (e.g., for MLST or cgMLST typing). Auto-correction of homopolymer related InDel errors (Ion PGM & 454 GS Junior).
Sanger data: Assemble, edit and analyze Sanger CE sequencing data (e.g., for 16S rDNA or MLST typing). Chromatograms can be automatically assembled into contigs using configurable file naming conventions.
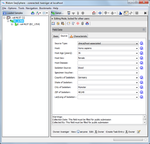
Database
Store, search, retrieve, export and create reports from your epidemiologic and DNA sequence data stored in an integrated SQL database.
Search new sequence entries against stored data. Data fields are compliant with the metadata requirements of the NCBI BioSample.
Data entry plausibility checks are possible.
Bacterial genotyping
Typing of bacteria is automatically performed. Ridom SeqSphere includes support for the following genotyping methods:
MLST: Fetch allele numbers for sequences and STs from the PubMLST server or define a local MLST typing, e.g. for using new targets.
cgMLST: Extension of MLST that uses thousands of targets, for a gene-by-gene allele typing of NGS data.
Automatic NCBI GenBank online queries: Compare the DNA sequences with best matches from GenBank.
Single locus queries: Do single locus typing (SLST) and compare against libraries stored in the local database.
Quality checking
Quality of DNA data can be automatically checked with user defined target QC parameters (e.g., coverage, intragenic stop codons, frame shifts detection).
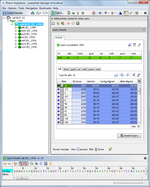
Analytical tools
Select data entries from a comparison table for epidemiologic, evolutionary or functional analysis. Cluster and visualize data entries by using minimum spanning or UPGMA/Neighbor Joining trees.
Security
Encryption (SSL) of data in transmission. Various configurable user roles, user groups and access controls. Audit trail functionality (who, when and what).
Community
Option to contribute to a single world-wide expanding publically available database of nomenclature and epidemiologic data. Own server for epidemiologic data for larger (supra) national institutions on request. Rapidly and easily share task templates with other institutions or download them online from the Task Template Sphere.