Introduction
MBioSEQ Ridom Typer implements GAMBIT (Genomic Approximation Method for Bacterial Identification and Tracking; citation), a bioinformatics tool with a curated database that uses genome assembly k‑mer comparisons to rapidly and accurately identify bacterial species. It uses taxon-specific predicted taxonomic rank estimations. GAMBIT does not do identification down to subspecies level and subsumes all Mycobacterium tuberculosis complex species as M. tuberculosis.
It works by hashing genome assemblies into targeted k‑mer signatures and comparing them against a reference database, but crucially it applies species‑specific thresholds to decide whether a match is reliable. Instead of using a single universal cutoff, GAMBIT determines for each species the minimum similarity score (based on shared k‑mers) that must be reached to confidently assign an isolate to that species. This design reduces false positives and accounts for natural genomic diversity across different taxa. As a result, GAMBIT can deliver rapid species identification with an accuracy comparable to average nucleotide identity (ANI) methods, but at a fraction of the computational cost. Furthermore, it is more discriminatory than ANI. Further details can be found in the documentation.
MBioSEQ Ridom Typer uses the Theiagen database v2.0.1 (citation) that is based on GTDB r214.1 (April 28th, 2023) as reference database for GAMBIT.
If a sample is processed in a pipeline and has a GAMBIT task entry, then the Genus and Species fields are derived from the Predicted Name (see Pipeline Script).
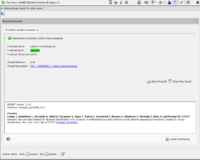
Task Entry Overview
The task entry overview shows the GAMBIT results for the sample.
Result Fields
The task entry stores the following result fields for each sample:
| Field | Description |
|---|---|
| Predicted Name | Taxon name assigned to the query (e.g., species or genus), based on GAMBIT's classification decision. |
| Predicted Rank | Taxonomic rank at which the prediction is made (e.g., species, genus). For QC this field is highlighted green if the rank is "species", else it is yellow. |
| Predicted Threshold | Decision threshold applied for the predicted rank; used to determine whether the match is confident at that rank. |
| Closest Distance | Distance score between the query and the closest reference genome in the GAMBIT database; lower means more similar. |
| Closest Description | Human-readable description of the closest reference (e.g., organism/strain and metadata). |
| Next Name | Taxon name of the next-best candidate considered by GAMBIT. |
| Next Rank | Taxonomic rank of the next-best candidate. |
| Next Threshold | Decision threshold relevant to the next-best candidate’s rank. |
The Result tab of the Sample Overview only shows the field Predicted Name. This field is also written to the Procedure Statistics. When a comparison table is created for a project that contains the GAMBIT Bacterial Species ID Task Template this field is automatically added to the comparison table, replacing the Top Species Match" from Mash. If the task template is explicitly selected when creating a comparison table, all result fields are added to the table.
Run times
| Species | NCBI ID | Run time |
|---|---|---|
| Escherichia coli | NC_000913.3 | 29 sec |
| Listeria monocytogenes | NC_003210.1 | 28 sec |
| Staphylococcus aureus | NZ_CP007455.1 | 28 sec |
FOR RESEARCH USE ONLY. NOT FOR USE IN CLINICAL DIAGNOSTIC PROCEDURES.