![]() Important: This page is only meant for Linux users. If you are using Windows and WSL, please see Windows Subsystem For Linux.
Important: This page is only meant for Linux users. If you are using Windows and WSL, please see Windows Subsystem For Linux.
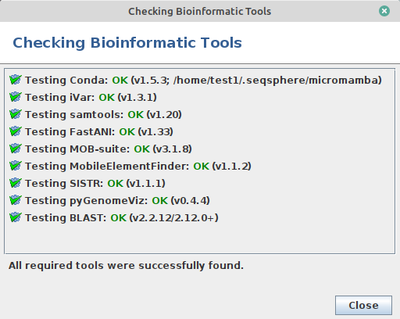
SeqSphere uses some bioinformatic tools that are not deployed together with the SeqSphere installation. Instead, they must be installed once for each Linux user: When SeqSphere is started for the first time, a check for those is performed automatically and a dialog is shown that allows to install missing tools. The check can also be invoked any time later by using the menu function Help | Check Bioinformatic Tools.
If the installation started, SeqSphere installs the missing tools as separate conda environments using micromamba into a subfolder of the profile directory (by default "~/.seqsphere/micromamba"). This does not conflict with any existing conda installation of the Linux user.
The following tools will be installed:
- iVar (SARS-CoV-2 tiled amplicon raw reads processing)
- SISTR (S. enterica geno-serotyping)
- MOB-suite (chromosome and plasmids detection)
- MobileElementFinder
- FastANI (whole genome average nucleotide identity)
- pyGenomeViz (genome visualization)
- Chopper (Long read preprocessing)1)
- Rasua (Long read preprocessing)1)
- FiltLong (Long read preprocessing)1)
- Flye (Long read assembling)1)
- Raven (Long read assembling)1)
- Medaka (Long read assembly polishing)1)
1) Only required and available for ONT Data Assembly Module
2) Only required and available for Long-read Data Plasmid Transmission Analysis Module
If no graphical user interface is available, the installation can also be invoked by command line parameter.
Updating from SeqSphere Version 9
SeqSphere version 9 used a Miniconda installation or any existing conda installation in the user's home to install the bioinformatic tools. With SeqSphere 10 this was changed to use separate micromamba envrionments that do not conflict with the user environment.
When updating from SeqSphere version 9, any existing conda installation that contains some of the tools is initially used by default. However, this conda installation can be used for the old functions, but does not contain the newest version of the tools and not all tools, that are required for some new features.
Therefore it is recommended to update the bioinformatic tool installation by
- Step 1: Uninstalling the old conda installation if it is not needed outside of SeqSohere (i.e., renaming or deleting the folder ~/.miniconda3 and removing the init block from ~/.bashrc).
- Step 2: Invoking the new micromamba based installation with Help | Check Bioinformatic Tools. If the old conda installation was not removed fully, it may be necessary to perform this installation twice until all tools are successfully installed and detected.